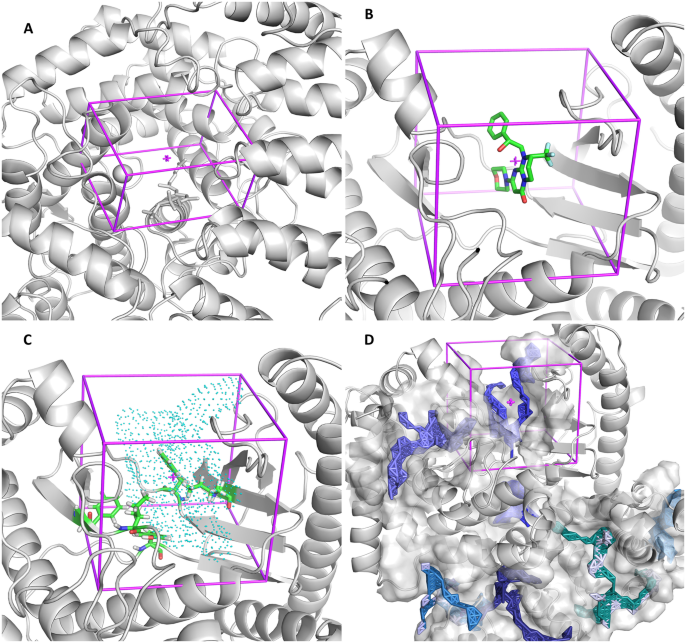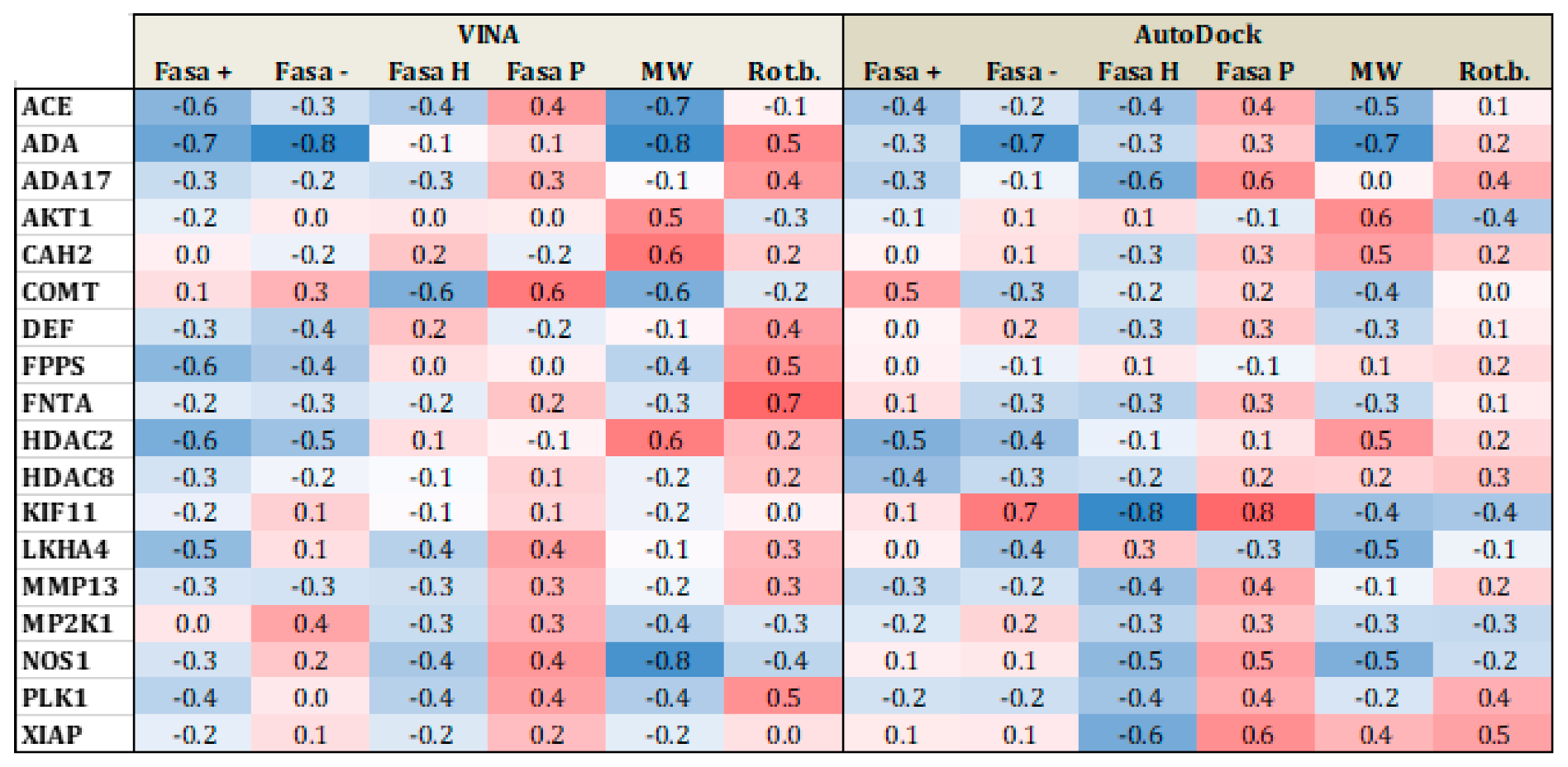
Applied Sciences | Free Full-Text | Comparing AutoDock and Vina in Ligand/Decoy Discrimination for Virtual Screening

AMDock: a versatile graphical tool for assisting molecular docking with Autodock Vina and Autodock4 | Biology Direct | Full Text

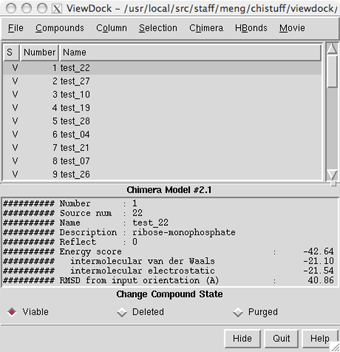
Identifying Protein-Ligand Interactions with Colab | by JOSÉ MANUEL NÁPOLES DUARTE | Towards Data Science

Evaluation of the binding performance of flavonoids to estrogen receptor alpha by Autodock, Autodock Vina and Surflex-Dock - ScienceDirect

AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading - Trott - 2010 - Journal of Computational Chemistry - Wiley Online Library

Molecular Docking Analysis | Autodock Results Analysis | Protein Ligand Int | Pymol | LigPlot Etc., - YouTube

Improving AutoDock Vina Using Random Forest: The Growing Accuracy of Binding Affinity Prediction by the Effective Exploitation of Larger Data Sets - Li - 2015 - Molecular Informatics - Wiley Online Library

Evaluation of AutoDock and AutoDock Vina on the CASF-2013 Benchmark | Journal of Chemical Information and Modeling

Webina: An Open-Source Library and Web App that Runs AutoDock Vina Entirely in the Web Browser | bioRxiv

AutoDock Vina 1.2.0: New Docking Methods, Expanded Force Field, and Python Bindings | Journal of Chemical Information and Modeling